Note
This tutorial was generated from a Jupyter notebook that can be downloaded here.
Inference on time-variable stars¶
In this tutorial we’ll show how to infer spot properties from a star whose spots evolve in time. Check out the ensemble tutorial for a bit more information on how we’re doing our analysis here.
Generate¶
Let’s generate a synthetic light curve of a star with the following properties:
[2]:
truths = dict(
r=15.0,
mu=30.0,
sigma=5.0,
c=0.05,
n=20,
tau=3.0,
p=1.123,
i=65.0,
u=[0.0, 0.0],
ferr=1e-3,
)
npts = 1500
tmax = 300.0
# Things we're solving for
varnames = ["r", "mu", "sigma", "c", "n", "i", "p", "tau"]
parameter |
description |
true value |
|---|---|---|
|
mean radius in degrees |
|
|
latitude distribution mode in degrees |
|
|
latitude distribution standard deviation in degrees |
|
|
fractional spot contrast |
|
|
number of spots |
|
|
stellar inclination in degrees |
|
|
stellar rotation period in days |
|
|
spot evolution timescale in days |
|
|
limb darkening coefficients |
|
|
photometric uncertainty (fractional) |
|
|
number of points in light curve |
|
|
length of light curve in days |
|
We will generate the light curve using a time-variable StarryProcess. If we were being rigorous about this, we should generate it from an actual forward model of a time-variable stellar surface. But that’s beyond the scope of this simple tutorial. Our goal here is to show that the light curve of a single star can encode most of the information we’re interested in if we observe it over many spot evolution timescales. This is something that cannot be done for a star with a static surface! The
time variability is actually giving us lots of information about the covariance in the modes we’re able to probe at the particular inclination we see the star at.
[4]:
from starry_process import StarryProcess
import numpy as np
t = np.linspace(0, tmax, npts)
sp = StarryProcess(**truths, marginalize_over_inclination=False)
flux0 = sp.sample(t, p=truths["p"], i=truths["i"], u=truths["u"]).eval()
flux = flux0 + truths["ferr"] * np.random.randn(*flux0.shape)
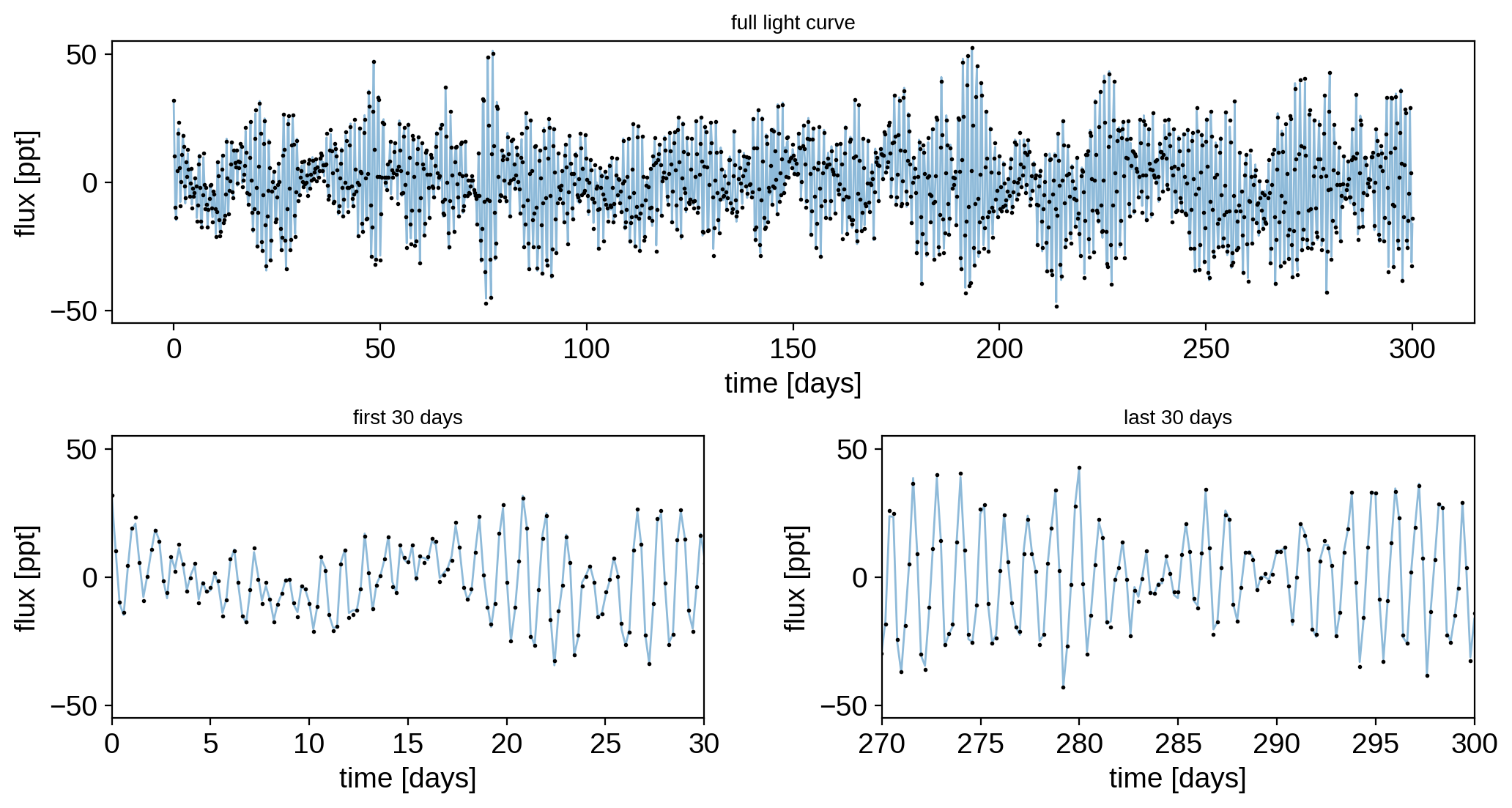
Here’s what our light curve looks like over all 300 days, as well as over the first and last 30 day segments:
[5]:
import matplotlib.pyplot as plt
fig = plt.figure(figsize=(12, 6))
fig.subplots_adjust(hspace=0.4, wspace=0.3)
ax = [
plt.subplot2grid((2, 2), (0, 0), colspan=2),
plt.subplot2grid((2, 2), (1, 0)),
plt.subplot2grid((2, 2), (1, 1)),
]
for axis in ax:
axis.plot(t, 1e3 * flux0[0], "C0-", lw=1, alpha=0.5)
axis.plot(t, 1e3 * flux[0], "k.", ms=2)
axis.set_ylabel("flux [ppt]")
axis.set_xlabel("time [days]")
axis.set_ylim(-55, 55)
ax[0].set_title("full light curve", fontsize=10)
ax[1].set_title("first 30 days", fontsize=10)
ax[2].set_title("last 30 days", fontsize=10)
ax[1].set_xlim(0, 30)
ax[2].set_xlim(270, 300)
plt.show()
Inference¶
Let’s set up a simple probabilistic model using pymc3 to infer the spot properties of this star. We’ll place uninformative priors on everything, including the period, timescale, and inclination. For simplicity, let’s assume we know the limb darkening coefficients (which are zero in this case).
Note
Note that we are passing marginalize_over_inclination=False when instatiating our
StarryProcess. That’s because we want to actually fit for the inclination of this
star. Marginalizing over the inclination is useful when we’re analyzing a large
ensemble of stars and don’t want to model the inclinations of every single one of
them (which usually makes sampling trickier and slower).
[6]:
import pymc3 as pm
import theano.tensor as tt
with pm.Model() as model:
# Things we know
u = truths["u"]
ferr = truths["ferr"]
# Spot latitude params. Isotropic prior on the mode
# and uniform prior on the standard deviation
unif0 = pm.Uniform("unif0", 0.0, 1.0)
mu = 90 - tt.arccos(unif0) * 180 / np.pi
pm.Deterministic("mu", mu)
sigma = pm.Uniform("sigma", 1.0, 20.0)
# Spot radius (uniform prior)
r = pm.Uniform("r", 10.0, 30.0)
# Spot contrast & number of spots (uniform prior)
c = pm.Uniform("c", 0.0, 0.5, testval=0.1)
n = pm.Uniform("n", 1.0, 30.0, testval=5)
# Inclination (isotropic prior)
unif1 = pm.Uniform("unif1", 0.0, 1.0)
i = tt.arccos(unif1) * 180 / np.pi
pm.Deterministic("i", i)
# Period (uniform prior)
p = pm.Uniform("p", 0.75, 1.25)
# Variability timescale (uniform prior)
tau = pm.Uniform("tau", 0.1, 10.0)
# Instantiate the GP
sp = StarryProcess(
r=r, mu=mu, sigma=sigma, c=c, n=n, tau=tau, marginalize_over_inclination=False
)
# Compute the log likelihood
lnlike = sp.log_likelihood(t, flux, ferr ** 2, p=p, i=i, u=u)
pm.Potential("lnlike", lnlike)
As we discussed in this tutorial, it can be very difficult to sample the posterior for a StarryProcess problem using Hamiltonian Monte Carlo. It’s often much easier to use regular old-fashioned MCMC, which we can easily due via the MCMCInterface class:
[7]:
from starry_process import MCMCInterface
with model:
mci = MCMCInterface()
Let’s run the optimizer to find the maximum a posteriori (MAP) solution:
[8]:
x = mci.optimize()
optimizing logp for variables: [tau, p, unif1, n, c, r, sigma, unif0]
message: Desired error not necessarily achieved due to precision loss.
logp: 2020.3826546101209 -> 5455.770611578508
Recall that we need to transform the solution from the internal parametrization to our desired parametrization (in terms of the spot radius, spot latitude, etc.):
[9]:
map_soln = mci.transform(x, varnames=varnames)
parameter |
description |
true value |
MAP value |
|---|---|---|---|
|
mean radius in degrees |
|
|
|
latitude distribution mode in degrees |
|
|
|
latitude distribution standard deviation in degrees |
|
|
|
fractional spot contrast |
|
|
|
number of spots |
|
|
|
stellar inclination in degrees |
|
|
|
stellar rotation period in days |
|
|
|
spot evolution timescale in days |
|
|
Not bad! It looks like we correctly estimated several of the properties of interest! We still don’t have any sense of the uncertainty on these quantities; for that, we need to actually run the MCMC sampler.
[11]:
nwalkers = 30
p0 = mci.get_initial_state(nwalkers)
Note
Since we’re running this example on GitHub Actions, we’ll only run the chain for 1000 steps, so it won’t be converged. You should run it for much, much longer if you can!
[12]:
import emcee
# Number of parameters
ndim = p0.shape[1]
# Instantiate the sampler
sampler = emcee.EnsembleSampler(nwalkers, ndim, mci.logp)
# Run the chains
np.random.seed(0)
nsteps = 1000
_ = sampler.run_mcmc(p0, nsteps, progress=True)
100%|██████████| 1000/1000 [1:33:43<00:00, 5.62s/it]
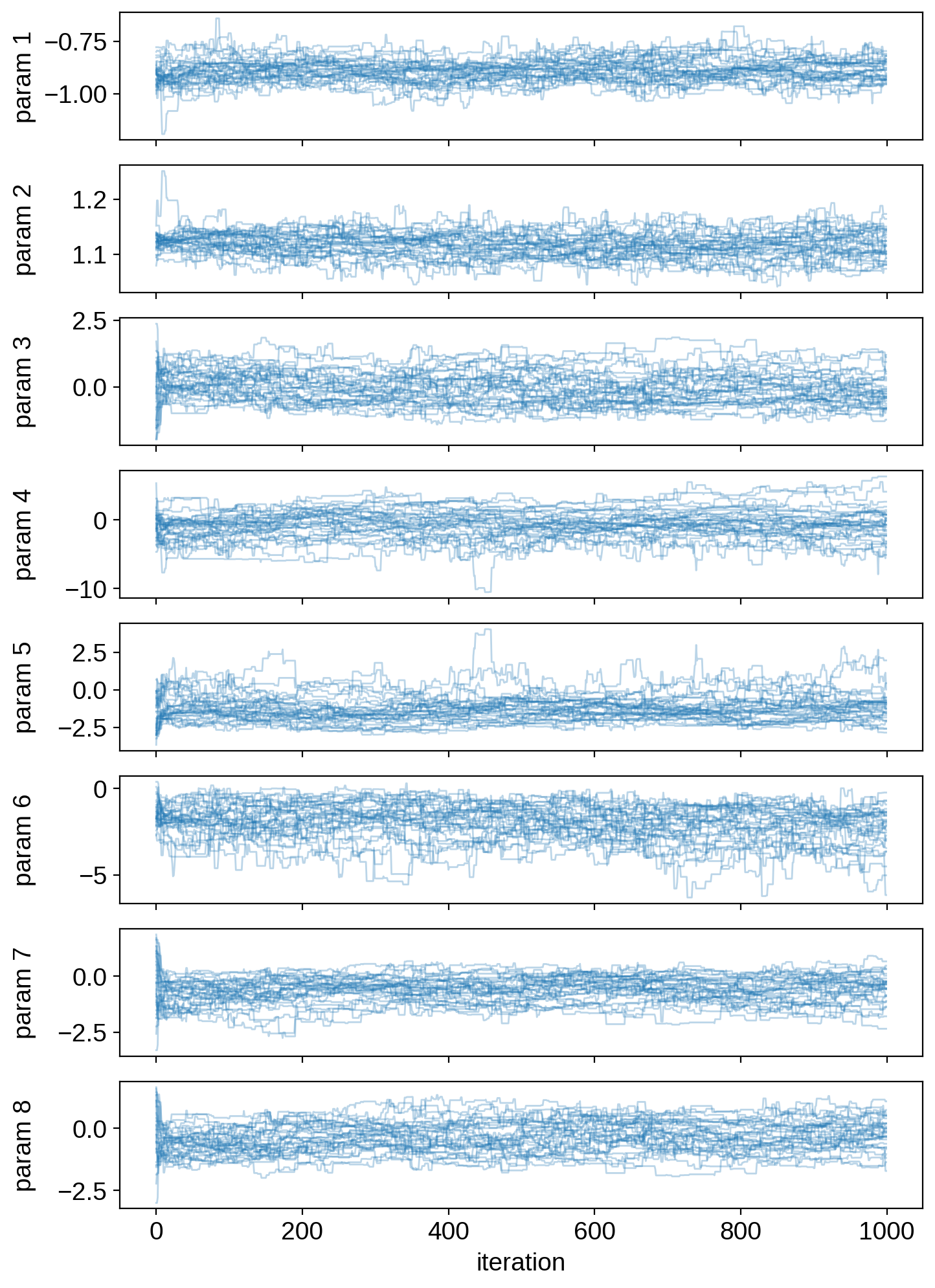
Let’s plot the chains:
[13]:
# Plot the walkers
fig, ax = plt.subplots(ndim, figsize=(8, 12), sharex=True)
for j in range(ndim):
for k in range(nwalkers):
ax[j].plot(sampler.chain[k, :, j], "C0-", lw=1, alpha=0.3)
ax[j].set_ylabel("param {}".format(j + 1))
ax[-1].set_xlabel("iteration")
fig.align_ylabels(ax)
plt.show()
Remove some burn-in and flatten the samples:
[14]:
burnin = 200
samples = sampler.chain[:, burnin:, :].reshape(-1, ndim)
Transform to our parametrization:
[15]:
samples = mci.transform(samples, varnames=varnames)
Visualize the posteriors, with the true values in orange:
[16]:
from corner import corner
corner(
samples,
labels=varnames,
truths=[truths[name] for name in varnames],
range=(
(10, 30),
(0, 90),
(1, 20),
(0, 0.5),
(1, 30),
(0, 90),
(0.75, 1.25),
(0.1, 10.0),
),
truth_color="C1",
bins=100,
smooth=2,
smooth1d=2,
)
plt.show()
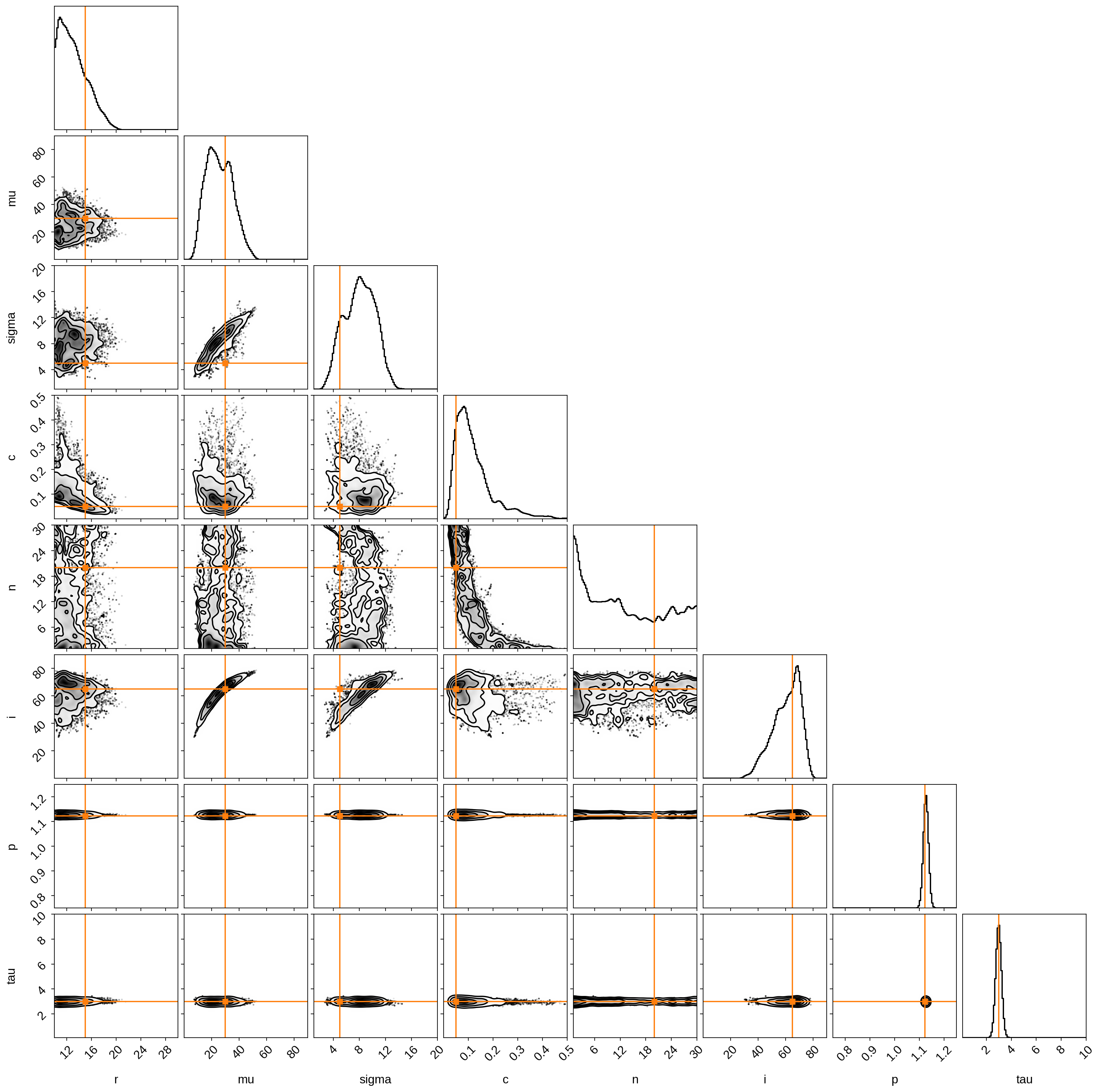
We did pretty good! Time-variable light curves encode a lot of the information we need, provided we observe the star over many (i.e., hundreds!) evolution timescales.